Overview
MITgcm.jl allows the analysis of MITgcm results in Julia. It can also setup, build, and launch a chosen model configuration (MITgcm_config) from within julia.
Functionalities are documented in the coming sections, and in Examples, Notebooks.
Getting Started
Installing the latest version of MITgcm.jl with the built-in package manager is the recommended method.
using Pkg
Pkg.add("MITgcm")
using MITgcm
system_check(setenv=true)The first time you do this, MITgcm.jl downloads a few MITgcm fortran code folders. This may take a few seconds to a few minutes depending on your network's performance.
Running MITgcm
Let's start running MITgcm interactively.
The first command defines a data structure, MITgcm_config, that we can then use from Julia. The setup command, by default, creates a folder in your tempdir() to run MITgcm.
using MITgcm
MC=MITgcm_config(configuration="tutorial_held_suarez_cs")
setup(MC)
show(MC) ID = 5dbcd567-674f-4c46-b75b-1ffe3845aa4f
model = MITgcm
configuration = tutorial_held_suarez_cs
run folder = /tmp/5dbcd567-674f-4c46-b75b-1ffe3845aa4f
log subfolder = /tmp/5dbcd567-674f-4c46-b75b-1ffe3845aa4f/log
task(s) = MITgcm_launchAfter setup, the standard workflow is to call build and then launch.
Next we can use the log method to get a status report.
build(MC)
launch(MC)
log(MC)3-element Vector{String}:
"17fcbf2 initial setup"
"642c96b initial tracked_parameters.toml"
"4d7e3b6 (HEAD -> main) add files in `tracked_parameters/` to git"If we have a previous build of MITgcm for this then we can specify the file path as a parameter, :exe.
If :exe is not specified, or if no file is found at the specified path, build will attempt to build the model for us.
exe=joinpath(default_path(),"verification",MC.configuration,"build","mitgcmuv")
MC.inputs[:setup][:build][:exe]=exe"/home/runner/.julia/scratchspaces/124859b0-ceae-595e-8997-d05f6a7a8dfe/datadeps/mitgcmsmall/MITgcm/verification/tutorial_held_suarez_cs/build/mitgcmuv"- For longer
MITgcmsimulations run, users often prefer to use a queuing system or batch script (not an interactive session). setupcan generate and submit a batch script, viacreate_scriptandconfig.inputs[:setup][:main][:command] = "qsub submit.csh"
Using Model Output
As MITgcm users, we often want to read and visualise model output from an earlier model run. To this end, MITgcm.jl provides methods to read the various file formats that MITgcm generates.
Read example:
T=read_mdsio(joinpath(MC,"run"),"T.0000276496")
size(T)(192, 32, 20)Plot example:
using CairoMakie
fig, ax, hm = heatmap(T[1:32,:,1])
ax.title="temperature"
Colorbar(fig[1, 2], hm)
fig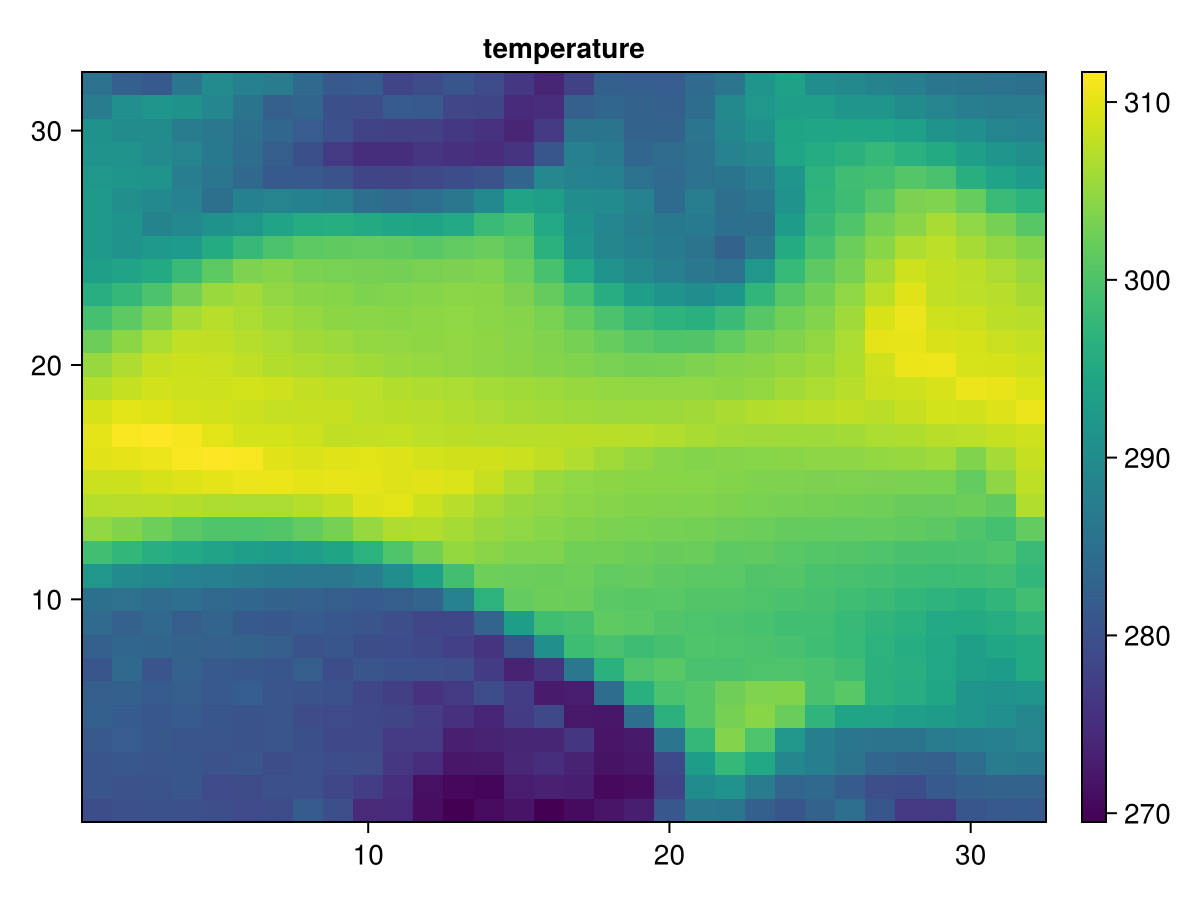
For more use cases, see MeshArrays.jl, Drifters.jl, Climatology.jl.
- geography notebook and vector tutorial present generic recipes, readily applicable to most MITgcm configurations
- ocean pathways can also be computed from MITgcm output
- climatology notebook shows a whole set of ocean variables, transports, etc computed from global MITgcm solutions (ECCO4 and OCCA2)
MITgcm File Formats
MITgcm stores model output within a run/ folder, such as the standard STDOUT text files, and other file formats listed below. scan_run_dir can be used to provide a summary of what's in the run/ folder. For more see:
- Standard Output (text)
- Input Files (text)
- MDS Files (binary output)
- MNC Files (netcdf output)
- Grid Files (binary or netcdf)
- Other Files
Grid variables are often needed for analysis. They can be read from file using either GridLoad_mdsio or GridLoad_mnc. This will return Γ.XC, Γ.YC, etc formated using MeshArrays.jl. See also GridLoad_native.
The MITgcm_scan_output.jl notebook does this in bulk for all configurations in MITgcm/verification and displays the gridded model domain for each model configuration (this page).
MITgcm Configurations
MITgcm.jl represents a model configuration using MITgcm_config. This data structure allows you take advantage of the ClimateModels.jl interface for example.
setupprepares a run directory for theMITgcm_configbuildcompiles the model (if needed)launchstarts the model run
The scan_verification function provides a list of standard model configurations. Each one has a subfolder in MITgcm_path[2] which gets compiled using source code from MITgcm_path[1].
For more on these aspects, see Examples, Model Configurations, and ClimateModels Interface.
Interactive notebooks can be found in the Examples section (and the examples/ subfolder). They demonstrate functionalities like plotting with Makie.jl and particle tracking with Drifters.jl.
Troubleshooting
The system_check method will try running MITgcm and report back. If the result is negative for any particular item, you may want to consult the MITgcm documentation for more guidance.
using MITgcm
MITgcm.system_check(setenv=true) MITgcm_system_check
name = advect_xy
download = true
complete = true
mpi = false
adj = false
folder = /tmp/850e0d76-0ff8-4ac9-8934-7acf708e48cc
path_MITgcm = /home/runner/.julia/scratchspaces/124859b0-ceae-595e-8997-d05f6a7a8dfe/datadeps/mitgcmsmall/MITgcm
path_verification = /home/runner/.julia/scratchspaces/124859b0-ceae-595e-8997-d05f6a7a8dfe/datadeps/mitgcmsmallverif/MITgcm/verification
NETCDF_ROOT = /usr
MPI_INC_DIR = /usr/lib/x86_64-linux-gnu/openmpi/include
genmake_log = OrderedDict with 5 keys
genmake_state = OrderedDict with 81 keys
MITgcm.setenv()can be used to setNETCDF_ROOTandMPI_INC_DIRto specified values.MITgcm.set_environment_variables_to_default()can be used to setNETCDF_ROOTandMPI_INC_DIRto default values.scan_build_dir,scan_run_dir, andmonitorcan be used to inspect the experiment folders.
- Building and running MITgcm requires a fortran compiler. Some configurations further require installing MPI and NetCDF libraries.
- The ECCO-Docker image has
MITgcm.jlpre-installed, as well asgfortran,MPI, andNetCDFallowing to run anyMITgcmconfiguration. The ECCO-Binder instance (free, but small) is available to try functionalities in the cloud.